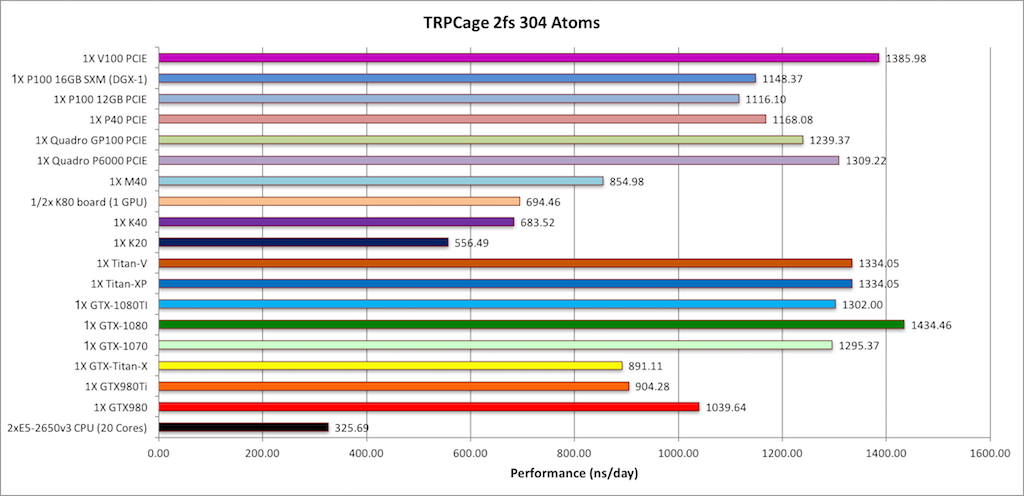
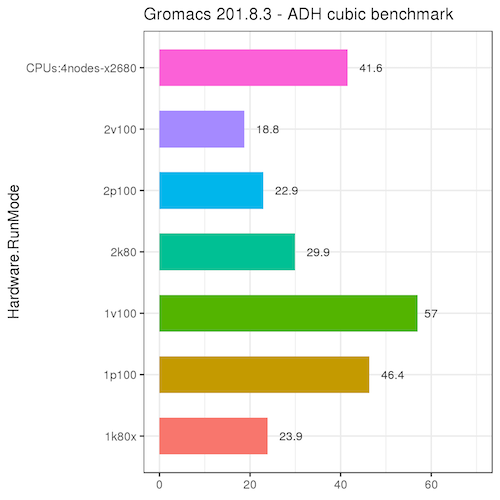
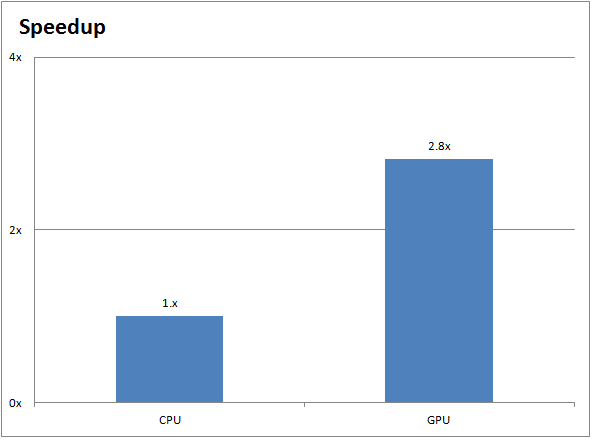Best bang for your buck: GPU nodes for GROMACS biomolecular simulations - Kutzner - 2015 - Journal of Computational Chemistry - Wiley Online Library

Comparison of job performance using GROMACS and ddCMD for one to four... | Download Scientific Diagram
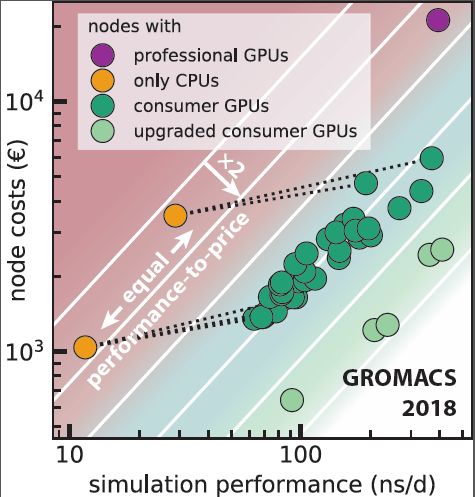
Webinar: More bang for your buck: Improved use of GPU Nodes for GROMACS 2018 (2019-09-05) – BioExcel – Centre of Excellence for Computation Biomolecular Research

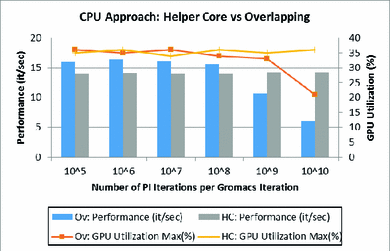
More bang for your buck: Improved use of GPU nodes for GROMACS 2018 - Kutzner - 2019 - Journal of Computational Chemistry - Wiley Online Library

Desmond, NAMD and GROMACS in comparison for standard benchmark protein... | Download Scientific Diagram